Research Team from the Laboratory: Metabolomic Analyses of Critically Ill COVID-19 Patients Unravel Biomarkers
2021-04-151275The First Affiliated Hospital of Guangzhou Medical University/State Key Laboratory of Respiratory Disease cooperated with BGI Research Institute, HKU-Pasteur Research Pole, the University of Hong Kong, the Sixth Affiliated Hospital of Guangzhou Medical University/Qingyuan People’s Hospital, the Fifth Affiliated Hospital of Sun Yat-sen University, Yangjiang People’s Hospital to analyze and compare the proteome, metabolome, and lipidome of serum and urine specimens from patients with mild and severe COVID-19. The study found prognostic markers and potential drug treatment targets for patients with severe COVID-19, and explained the mechanism of SARS-CoV-2 infection based on multi-omics data, providing theoretical basis and support for the analysis of the pathogenic mechanism of SARS-CoV-2 and the screening of COVID-19 therapeutic drugs. The research results were published online in Signal Transduction and Targeted Therapy, a sub-Journal of Nature, on April 15, 2021.

Since the outbreak of the COVID-19 epidemic a year ago, the number of infections has continued to rise. Currently it has infected more than 110 million people and resulted in more than 2.5 million deaths worldwide. Although the overall mortality rate for COVID-19 patients is 2.1%, the mortality rate for critically ill patients is as high as 20%. As a result, it is particularly important to predict the COVID-19 severity in advance to reduce mortality. This study conducted multi-omics and random forest analysis on plasma and urine specimens of patients with mild and severe COVID-19, and found that the combination of 4 proteins and 21 metabolites can accurately distinguish patients with mild COVID-19 from those with severe COVID-19. Using these molecules to verify the urine samples of patients, and we find that the accuracy rate of urine sample verification is as high as 90%. This conclusion made it possible for non-invasive sampling to monitor disease progression.
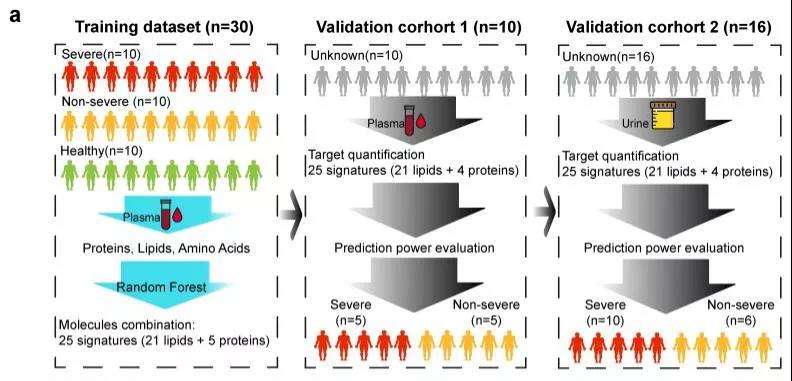
Experimental design and process
The research team also found that there are multiple different molecules in COVID-19 patients that can be used as potential therapeutic targets, some corresponding therapeutic drugs have been put into clinical trials, such as dexamethasone. Moreover, the research data also reveals the changes of molecules involved in macrophage polarization, T cell exhaustion and inflammatory response after infected with COVID-19. In the end, the research team combined the patient’s clinical manifestations and research related to cellular immunity to deduct the pathogenic mechanism of SARS-CoV-2, providing an important reference for monitoring the disease progression and adjusting the treatment plan.
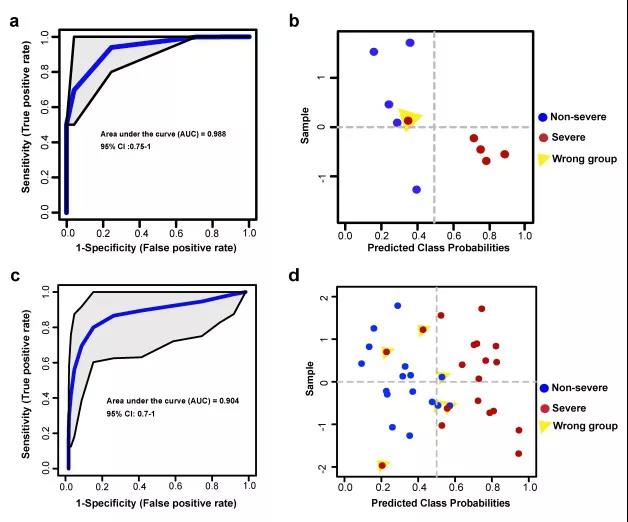
Predict COVID-19 severity through plasma (A and B) and urine (C and D) specimens
















